BenchDrop-seq platform
The Srivastava Lab builds BenchDrop-seq, a microfluidics-free benchtop platform that combines PIP-seq single-cell capture with Oxford Nanopore long-read sequencing. It runs on any standard lab bench without a dedicated instrument, puts isoform-level single-cell transcriptomics within reach of any group with a thermocycler, and extends naturally to chromatin accessibility with PIP-ATAC. Every protocol is open, reproducible, and engineered to scale from pilot experiments to atlas-scale immune profiling.
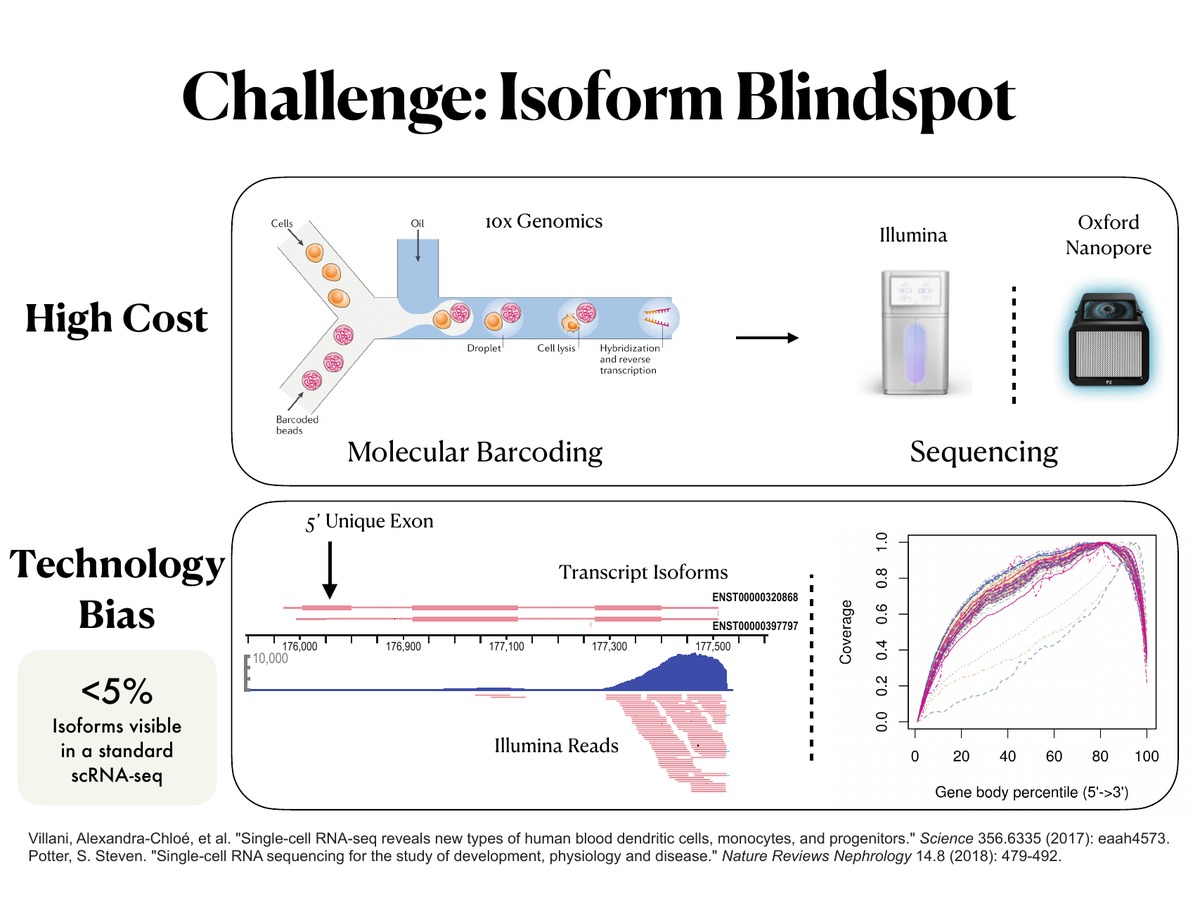
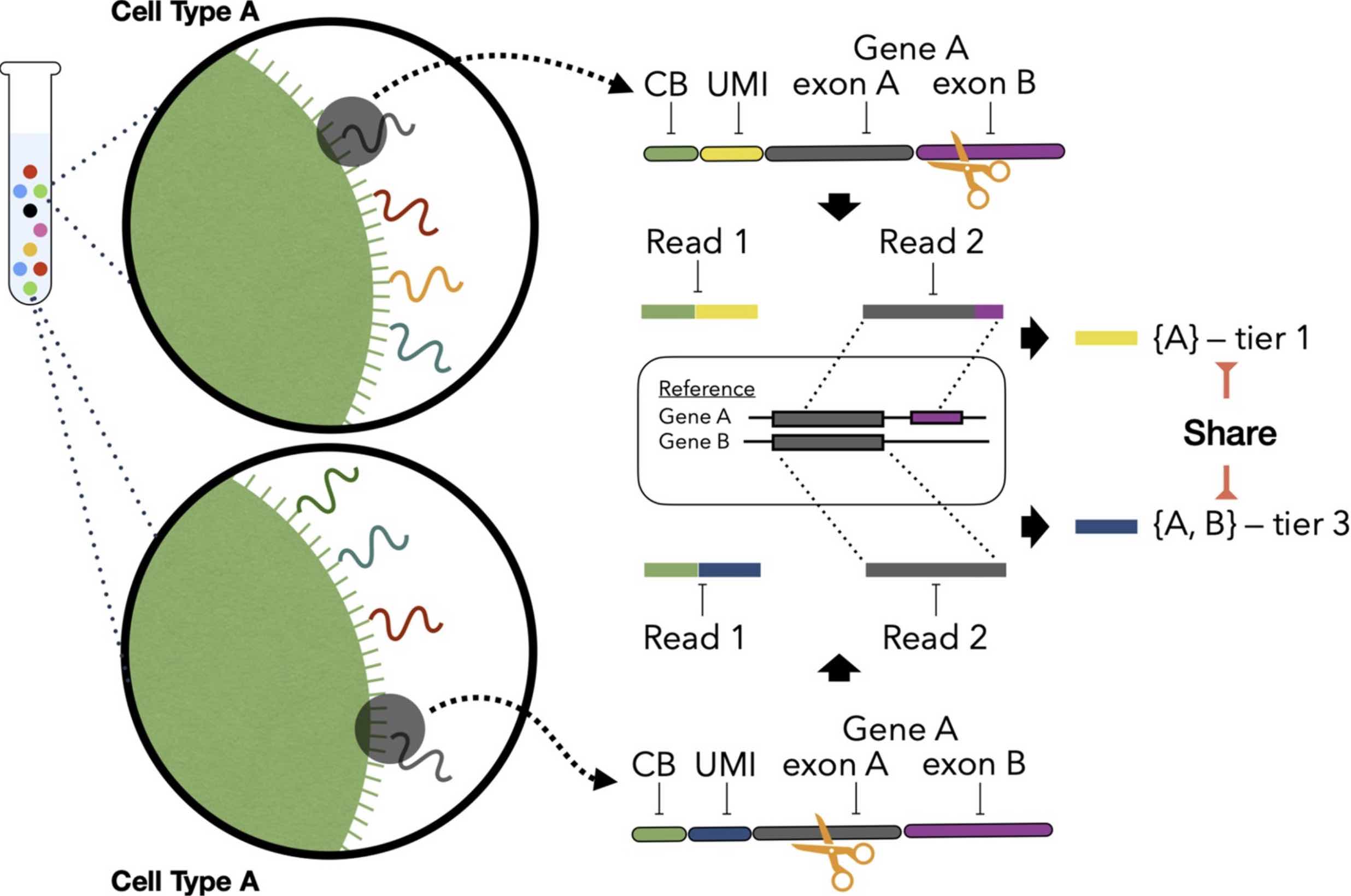
Bagpiper isoform pipeline
Long-read single-cell data is computationally distinct from short-read data: barcodes drift, reads disagree about which isoform they came from, and UMI deduplication has to honour splice-junction evidence rather than 3′ anchor alone. Bagpiper handles these problems end-to-end: spacer-anchored barcode recovery, junction-based read assignment, expectation-maximization multi-mapping resolution, and principled isoform × cell count matrices. The result is a reproducible path from raw Nanopore signal to isoform-resolved quantification that downstream analyses can actually trust.
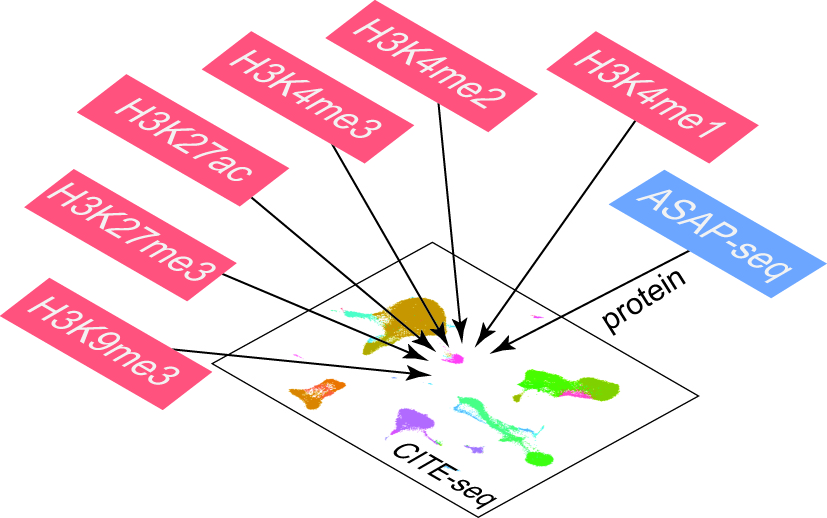
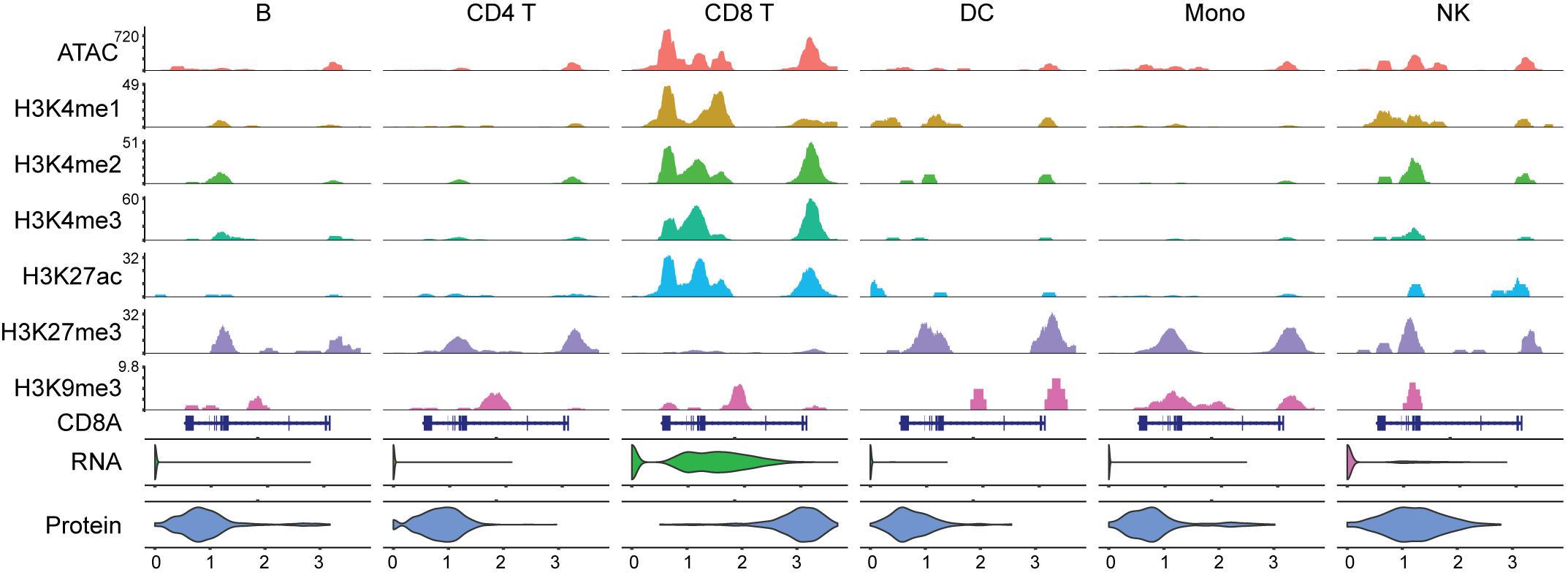
Cross-platform projection & multimodal integration
Isoform-level measurements only become interpretable when they sit alongside gene-level atlases, chromatin state, and surface-protein markers. The lab builds two-layer annotation strategies that transfer cell identity from established short-read references onto long-read BenchDrop-seq data, and multimodal integration frameworks that align transcriptome, chromatin, and isoform information into a single coherent state per cell. These methods make BenchDrop-seq datasets immediately comparable to existing immune and tumor atlases, without forcing users to choose between depth and breadth.
Cancer and autoimmunity, as co-equal programs
The lab runs two biological programs side by side rather than ranking one over the other. In cancer, BenchDrop-seq is applied to NRAS-mutant melanoma to identify the drug-tolerant persister states that survive pan-RAS inhibition and to track the RRAS isoform switching that accompanies resistance, in collaboration with the Villanueva Lab at Wistar.
In autoimmunity, the same pipeline profiles multiple sclerosis patient immune cells captured with Parse Biosciences and EBV-driven immune dysregulation in ex vivo PBMC models, asking where splicing rewires T cell and B cell fate decisions — in collaboration with the Lieberman Lab at Wistar. Cancer and autoimmunity are treated as two windows onto the same underlying question: how post-transcriptional regulation decides cell state.