Understand your cells,
layer by layer
Matisse brings isoform-resolved splicing and chromatin accessibility into your single-cell workflow — on top of Seurat and Signac, using the same cells, the same clusters, the same UMAP.
What you can discover
Questions Matisse is built to answer
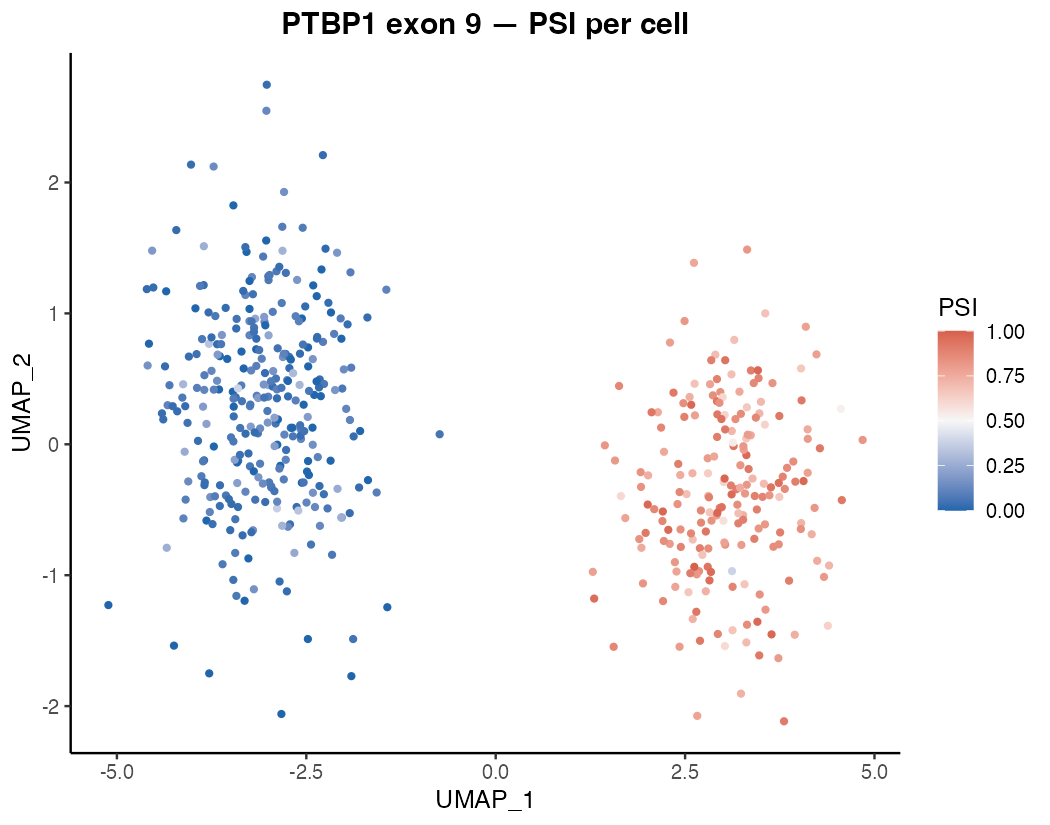
Cell-type-specific splicing
Do my neurons and astrocytes process this exon differently — and by how much?
Chromatin shapes isoforms
Is the splicing switch I see linked to chromatin accessibility changes at the same locus?
Isoform switches along a trajectory
Is there a coordinated splicing change as my cells differentiate or respond to a stimulus?
Context for bulk RNA-seq
I see a splicing difference in bulk data — which cell type is responsible?
Works with your existing setup
Matisse layers on top of Seurat and Signac — your clusters, UMAP, and cell labels stay intact
Short-read RNA (10x) STAR / STARsolo junction count matrix
Long-read / isoform Bagpiper FLAMES LIQA PacBio MAS-seq
ATAC / chromatin 10x Multiome Signac ArchR
Event annotations SUPPA2 generateEvents rMATS BuildSimpleEvents()
Installation
install.packages("remotes")
remotes::install_github("avisrilab/Matisse")